Chimera: Difference between revisions
No edit summary |
No edit summary |
||
| Line 8: | Line 8: | ||
Here are my general use-cases for Chimera (illustrated with the PDB-file 5ceh). If the command line is used no '|' character is used. The commandline can be enabled Header | Favorites | Command Line, which would indicate in the screen's header click Favorites and then Command Line. | Here are my general use-cases for Chimera (illustrated with the PDB-file 5ceh). If the command line is used no '|' character is used. The commandline can be enabled Header | Favorites | Command Line, which would indicate in the screen's header click Favorites and then Command Line. | ||
[[File:Binding 5ceh.png|400px|thumb|left| | [[File:Binding 5ceh.png|400px|thumb|left|Protein-ligand interactions in PDB ID 5ceh. Ligand is displayed in cyan, protein in brown, the metal co-factor in green.]] | ||
=== Showing the protein-ligand interactions === | === Showing the protein-ligand interactions === | ||
''open 5ceh'', loads 5ceh.pdb from the RCSB.<br /> | ''open 5ceh'', loads 5ceh.pdb from the RCSB.<br /> | ||
| Line 24: | Line 24: | ||
Click the 'Contact' button.<br /> | Click the 'Contact' button.<br /> | ||
Click apply.<br /> | Click apply.<br /> | ||
This will give you something like this | This will give you something like the image at the top of this section (the purple contact lines are hard to see with the black background, this can be fixed with Header | Actions | Color | all options and then selecting background and clicking white). | ||
Revision as of 15:21, 28 June 2024
Chimera is a molecular visualization program developed at UCSF (https://www.cgl.ucsf.edu/chimera/). It can be obtained for free for non-commercial use and can also be easily installed on your own computer. Chimera is a Python-based program.
General tips for usage
Most Chimera operations can be performed with commands via the commandline, by clicking and/or by using the toolbar in the top. I would recommend following the Tutorials in the Help section for first-time users (Header | Help | Tutorials). In general the program functions by making a selection of particular set of atoms (for instance a ligand) and then performing some action on them. If the action is more advanced than colouring or representation of the atoms, you can generally find it in Header | Tools. Each file you load in will be loaded as a separate model (shown at the bottom of the screen). The file you load in first will be #0, the second file #1 etc. Keep in mind that if a file contains multiple molecules (for instance a combined .mol2 file), the molecules are referenced individually as #0.1 #0.2 etc.
It is also important to realize a PDB-file can contain multiple molecules. When these are multiple protein molecules, these are stored in separate so-called chains. If you are in doubt what is stored in what chain, use the info on the PDB's page on the RCSB.
Here are my general use-cases for Chimera (illustrated with the PDB-file 5ceh). If the command line is used no '|' character is used. The commandline can be enabled Header | Favorites | Command Line, which would indicate in the screen's header click Favorites and then Command Line.
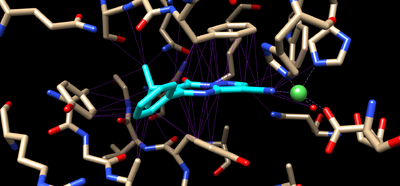
Showing the protein-ligand interactions
open 5ceh, loads 5ceh.pdb from the RCSB.
sel ligand, selects all atoms recognized as the ligand in the pdb-file.
col cyan sel, colors the selected atoms cyan.
col byhet sel, colors the selected atoms by heteroatom (the carbons will retain the original color; these steps could have been done at once with col cyan sel & C which is a conditional).
~disp, hide all default displayed atoms.
disp #0 & sel z < 6, shows all residues within 6 Ångstrom of the selection.
~ribbon, hides the cartoon representation of helices and sheets.
focus sel, set the selection at the center of the screen and hide far-away atoms.
Header | Tools | Surface/Binding Analysis | Find Clashes/Contacts.
In the pop-up window 'Find Clashes/Contacts' with the ligand still selected click 'Designate selection as second set'.
Sel ~sel, select everything that wasn't selected.
Click in the same window 'Designate' currently selected atoms for checking.
Click the 'Contact' button.
Click apply.
This will give you something like the image at the top of this section (the purple contact lines are hard to see with the black background, this can be fixed with Header | Actions | Color | all options and then selecting background and clicking white).
Tips for cluster usage
Chimera should generally only be used on the cluster if you want to run a large-scale experiment (for instance downloading 500 PDB files and superposing them). In such cases it can be useful to either:
- Run Chimera without a graphical user interface (GUI) with chimera --nogui
- Use X11 forwarding to forward the cluster screen. This requires upon each execution of the ssh command that you add the -X -Y flags which would result in ssh -X -Y user@cn142.science.ru.nl. In the case of Windows Powershell users, this may not function correctly due to antivirus scanners/firewalls. In such case, using the windows subsystem linux (WSL) can offer a solution.